Bayesian Learning
1 Introduction
- Bayesian learning methods provide useful learning
algorithms and help us understand other learning
algorithms.
- The practical learning algorithms are:
- Naive Bayes learning.
- Bayesian belief network learning—combines prior knowledge with observed data.
- Bayes reasoning provides the "gold standard" for evaluating other algorithms.
2 Bayes Theorem
- Bayes theorem [3] states that
\[ P(h\,|\,D) = \frac{P(D\,|\,h) P(h)}{P(D)} \]
where
- $P(h)$ = prior probability of hypothesis $h$
- $P(D)$ = prior probability of training data $D$
- $P(h\,|\,D)$ = probability of $h$ given $D$, also known as the posterior probability of $h$.
- $P(D\,|\,h)$ = probability of $D$ given $h$
2.1 Choosing a Hypothesis
- Generally want the most probable hypothesis given the
training data.
- The maximum a posteriori hypothesis is $h_{MAP}$ where
\[
\array{
h_{MAP} & = \arg \max_{h \in H} P(h\,|\,D) \\
& = \arg \max_{h \in H} \frac{P(D\,|\,h) P(h)}{P(D)} \\
& = \arg \max_{h \in H}P(D\,|\,h) P(h)
} \] we dropped $P(D)$ because its independent of $h$.
- If we assume that all hypothesis have the same prior
probability, that is $\forall_{i \neq j}P(h_i) = P(h_j)$, then
we can simplify even more and choose the maximum likelihood hypothesis
\[h_{ML} = \arg \max_{h \in H} P(D\,|\,h) \]
2.2 Example
- A patient takes a lab test and the result comes back positive. The test
returns a correct positive result in only $98%$ of the
cases in which the disease is actually present, and a
correct negative result in only $97%$ of the cases in which
the disease is not present. Furthermore, $.008$ of the
entire population have this cancer.
\[ \array{ P(cancer) = .008 & P(\neg cancer) = .992 \\ P(\oplus \,|\,
cancer) = .98 & P(\ominus \,|\, cancer) = .02 \\ P(\oplus \,|\, \neg cancer) =
.03 & P(\ominus \,|\, \neg cancer) = .97} \]
- If a new patient comes in with a positive test result,
whats the probability that he has cancer?
\[
\array{
P(\oplus\,|\,cancer) P(cancer) = (.98).008 = .0078 \\
P(\oplus\,|\,\neg cancer) P(\neg cancer) = (.03).992 = .0298
}
\] so $h_{MAP} = \neg cancer$.
- Product rule: probability $P(A \wedge B)$ of a conjunction of two events
A and B: \[P(A \wedge B) = P(A\,|\,B) P(B) = P(B\,|\,A) P(A) \]
- Sum rule: probability of a disjunction of two events A and B:
\[P(A \vee B) = P(A) + P(B) - P(A \wedge B) \]
- Theorem of total probability: if events $A_{1}, \ldots, A_{n}$ are mutually exclusive with $\sum_{i = 1}^{n} P(A_{i}) = 1$, then
\[P(B) = \sum_{i = 1}^{n} P(B\,|\,A_{i}) P(A_{i})\]
3 Brute Force Bayes Concept Learning
- Brute-Force MAP Learning algorithm:
- For each hypothesis $h$ in $H$, calculate the posterior probability
\[ P(h\,|\,D) = \frac{P(D\,|\,h) P(h)}{P(D)}\]
where $D = (d_1\ldots d_m)$ is the set of target values from the set of
examples $X = ((x_1,d_1) \cdots (x_m,d_m))$.
- Output the hypothesis $h_{MAP}$ with the highest posterior probability
\[h_{MAP} = \argmax_{h \in H} P(h\,|\,D)\]
- This can be very slow since it applies BT to every $h\in H$.
3.1 Relation to Concept Learning
- In concept learning we have an instance space $X$,
hypothesis space $H$, training examples $D$. The
FindS algorithm finds the most specific hypothesis
from $VS_{H,D}$.
- Assume fixed set of instances $\langle x_{1}, \ldots, x_{m}\rangle$
- Assume $D$ is the set of classifications $D = \langle c(x_{1}),
\ldots, c(x_{m})\rangle$
- Choose $P(D\,|\,h)$
\[
P(D\,|\,h) = { \{
\array{
1 & if \forall_{d_i \in D} d_i = h(x_i)\\
0 & otherwise
} }
\]
- Choose $P(h)$ to be uniform distribution so that $P(h) =
\frac{1}{|H|}$ for all $h$ in $H$
- Then we can say that
\[ P(h\,|\,D) = { \{ \array{ \frac{1}{|VS_{H,D}|} & if h is consistend
with D \\ 0 & otherwise. } } \]
- What happens in Brute force
BCL is that all hypotheses start with with same
probability then the inconsistent hypotheses drop to zero
while the rest share equally.
- Every consistent hypothesis is a MAP hypothesis.
3.2 MAP Hypothesis and Consistent Learners
- We showed that, in the given setting, every hypothesis
consistent with $D$ is a MAP hypothesis.
- Let a consistent learner be one that outputs a
hypothesis with zero error over training examples.
- Every consistent learner outputs a MAP hypothesis
if we assume uniform prior over $H$ and deterministic
noise-free examples.
- For example, since Find-S outputs a consistent
hypothesis, it outputs a MAP hypothesis, all without
manipulating probabilities!
4 Learning A Real-Valued Function
- Consider any real-valued target function $f$.
- Training examples $\langle x_{i}, d_{i} \rangle$, where $d_{i}$ is noisy
training value. $d_{i} = f(x_{i}) + e_{i}$, and $e_{i}$ is random
variable (noise) drawn independently for each $x_{i}$ according to
some Gaussian distribution with mean=0.
- The maximum likelihood hypothesis $h_{ML}$ we defined earlier as
\[
\array{
h_{ML} &= \argmax_{h \in H} p(D\,|\,h) \\
&= \argmax_{h \in H} \prod_{i=1}^{m} p(d_{i}\,|\,h)
}
\]
if we replace that probability with its equation and simplify we will eventually get
\[
h_{ML} = \argmin_{h \in H} \sum_{i=1}^{m} \left(d_{i} -
h(x_{i})\right)^{2} \]
- Therefore, $h_{ML}$ is the hypothesis that minimizes the
sum of the squared errors, if observations are generated by
adding Normal noise with zero main to the true data.
- Under these conditions any learning algorithm that minimizes the squared error will output a maximum likelihood hypothesis.
5 Learning To Predict Probabilities
- Consider predicting survival probability from patient data
- Training examples $\langle x_{i}, d_{i} \rangle$, where
$d_{i}$ is 1 or 0
- Want to train neural network to output a probability given
$x_i$ (not a 0 or 1)
- In this case can show
\[ h_{ML} = \argmax_{h \in H} \sum_{i=1}^{m} d_{i} \ln h(x_{i}) + (1-d_{i})
\ln (1 - h(x_{i})) \]
The negation of this quantity is known as the cross entropy.
- In order to maximize that we would need to do gradient
ascent on it, wrt the edge weight. This weight update works
out to be
\[ w_{jk} \leftarrow w_{jk} + \Delta w_{jk}\]
where
\[ \Delta w_{jk} = \eta \sum_{i=1}^{m} (d_{i} - h(x_{i})) x_{ijk} \]
- This is the same rule used by Backpropagation except that
Backpropagation multiplies by an extra term $h(x_i)(1 -
h(x_i))$, which is the derivative of the sigmoid
function.
- Backpropagation updates seek ML hypothesis under the
assumption that training data can be modeled by Normal noise
on the target function.
- Cross entropy updates seek ML hypothesis under the
assumption that observed boolean value is a probabilistic
function of input instance.
6 Minimum Description Length Principle
- We can interpret the definition of $h_{MAP}$ in terms of the basic concepts of information theory
\[
\array{
h_{MAP} &= &\arg \max_{h \in H}P(D\,|\,h) P(h) \\
&= &\arg \max_{h \in H} \log_{2} P(D\,|\,h) + \log_{2} P(h) \\
&= &\arg \min_{h \in H} - \log_{2} P(D\,|\,h) - \log_{2} P(h)
}
\]
This equation can be interpreted as a statement that short hypotheses
are preferred!
- $- \log_{2} P(h)$ is length of $h$ under optimal
code
- $- \log_{2} P(D\,|\,h)$ is length of $D$ given $h$ under optimal code
- The minimum description length principle
recomends choosing the hypothesis that minimizes the sum of
these two description lengths.
- If a representation of hypotheis is chosen so that the
size of $h$ is $- \log_2 P(h)$ and if a representation for
exceptions is chosen so that the encoding length of $D$ given
$h$ is $- \log_2 P(D\,|\,h)$ then the MDL principle produces MAP
hypotheses.
7 Bayes Optimal Classifier
- So far we've sought the most probable hypothesis given the
data $D$ (i.e., $h_{MAP}$)
- Given new instance $x$, what is its most probable
classification?
- $h_{MAP}(x)$ is not the most probable classification!
- For example, given three possible hypotheses:
\[
P(h_{1}\,|\,D)=.4, P(h_{2}\,|\,D)=.3, P(h_{3}\,|\,D)=.3
\]
($h_1$ is the MAP hypothesis) and given a new instance $x$,
\[
h_{1}(x)=\oplus, h_{2}(x)=\ominus, h_{3}(x)=\ominus
\]
what is the most probable classification of $x$, $\oplus$ or $\ominus$?
- The MAP hypothesis ($h_1$) says it is $\oplus$, but if
we consider all hypothesis they say that is is $\ominus$
with probability .6. So the most probable classification is
$\ominus$.
7.1 Bayes Optimal Classification
- Bayes optimal classification
\[ \arg \max_{v_{j} \in V} \sum_{h_{i} \in H} P(v_{j}\,|\,h_{i}) P(h_{i}\,|\,D)\]
- Example:
\[ \array{
P(h_{1}\,|\,D)=.4, & P(\ominus \,|\,h_{1})=0, & P(\oplus \,|\,h_{1})=1 \\
P(h_{2}\,|\,D)=.3, & P(\ominus \,|\,h_{2})=1, & P(\oplus \,|\,h_{2})=0 \\
P(h_{3}\,|\,D)=.3, & P(\ominus \,|\,h_{3})=1, & P(\oplus \,|\,h_{3})=0
} \]
therefore
\[ \array{
\sum_{h_{i} \in H} P(\oplus \,|\,h_{i}) P(h_{i}\,|\,D) & = & .4 \\
\sum_{h_{i} \in H} P(\ominus \,|\,h_{i}) P(h_{i}\,|\,D) & = & .6
} \]
and
\[ \array{
\arg \max_{v_{j} \in V} \sum_{h_{i} \in H} P(v_{j}\,|\,h_{i}) P(h_{i}\,|\,D) & = & \ominus
}\]
8 Gibbs Algorithm
- Bayes optimal classifier provides best result, but can be expensive if many
hypotheses.
- Gibbs algorithm:
- Choose one hypothesis at random, according to
$P(h\,|\,D)$
- Use this to classify new instance
- Surprising fact: Assume target concepts are drawn at
random from $H$ according to priors on $H$. Then:
\[ E[error_{Gibbs}] \leq 2 E[error_{Bayes Optimal}] \]
So, if the learner correctly assumes a uniform prior distribution over $H$, then
- Pick any hypothesis from VS, with uniform
probability
- Its expected error is no worse than twice Bayes optimal
9 Naive Bayes Classifier
- The naive bayes classifier is among the set of practical
learning methods.
- It should be used when
- There is large set of training examples.
- The attributes that describe instances are
conditionally independent given classification.
- It has been used in many applications such as diagnosis
and the classification of text documents.
9.1 Naive Bayes Classifier
- Assumes all instances $x \in X$ are described by attributes $\langle a_{1}, a_{2} \ldots a_{n} \rangle$.
- Assume target function $f: X \rightarrow V$, where $V$ is some finite set.
- Most probable value of $f(x)$ given $x = \langle a_{1}, a_{2} \ldots a_{n} \rangle$ is:
\[ \array{
v_{MAP} &= &\argmax_{v_{j} \in V} P(v_{j} \,|\, a_{1}, a_{2} \ldots a_{n}) \\
v_{MAP} &= &\argmax_{v_{j} \in V} \frac{P(a_{1}, a_{2} \ldots a_{n}\,|\,v_{j})
P(v_{j})}{P(a_{1}, a_{2} \ldots a_{n})} \\
v_{MAP} &= &\argmax_{v_{j} \in V} P(a_{1}, a_{2} \ldots a_{n}\,|\,v_{j}) P(v_{j})
} \]
- Naive Bayes assumes conditional independence given the target value, that is
\[ P(a_{1}, a_{2} \ldots a_{n}\,|\,v_{j}) = \prod_{i} P(a_{i} \,|\, v_{j}) \]
- Naive Bayes classifier:
\[ v_{NB} = \argmax_{v_{j} \in V} P(v_{j}) \prod_{i} P(a_{i} \,|\, v_{j})
\]
9.2 Naive Bayes Algorithm
- Naive Bayes Algorithm has a learning and a classifying functions.
- Naive_Bayes_Learn(examples)
- For each target value $v_j$
- $\hat{P}(v_j) \leftarrow $ estimate $P(v_j)$
- For each attribute value $a_i$ of each attribute $a$ $\hat{P}(a_i\,|\,v_j) \leftarrow$ estimate $P(a_i\,|\,v_j)$
- Classify_New_Instance($x$)
\[ v_{NB} = \argmax_{v_{j} \in V} \hat{P}(v_{j}) \prod_{a_i \in x} \hat{P}(a_{i} \,|\, v_{j}) \]
9.3 Naive Bayes Example
- We are trying to determine if its time to PlayTennis given that
\[\langle Outlook=sunny, Temp=cool, Humid=high, Wind=strong \rangle \]
- We want to compute
\[ v_{NB} = \argmax_{v_{j} \in V} P(v_{j}) \prod_{i} P(a_{i} \,|\, v_{j}) \]
- Given this set of examples we can calculate that
\[ P(PlayTennis = yes) = \frac{9}{14} = .64 \]
\[ P(PlayTennis = no) = \frac{5}{14} = .36 \]
similarly
\[P(Wind = strong \,|\, PlayTennis = yes) = \frac{3}{9} = .33 \]
\[P(Wind = strong \,|\, PlayTennis = no) = \frac{3}{5} = .6 \]
using these and few more similar probabilities we can calculate
\[P(yes) \cdot P(Outlook=sunny \,|\, yes)\cdot P(Temperature = cool \,|\, yes)\cdot P(Humidity = high \,|\, yes)\cdot P(Wind = strong \,|\, yes) = .005 \]
\[P(no)\cdot P(Outlook=sunny \,|\, no)\cdot P(Temperature = cool \,|\, no)\cdot P(Humidity = high \,|\, no)\cdot P(Wind = strong \,|\, no) = .021 \]
- So
\[ v_{NB} = no \]
9.4 Naive Bayes Issues
- Conditional independence assumption is often violated
\[ P(a_{1}, a_{2} \ldots a_{n}\,|\,v_{j}) = \prod_{i} P(a_{i} \,|\, v_{j}) \]
but it works surprisingly well anyway.
- We don't need estimated
posteriors $\hat{P}(v_j\,|\,x)$ to be correct; need only that
\[\argmax_{v_{j} \in V} \hat{P}(v_{j}) \prod_{i} \hat{P}(a_{i} \,|\, v_{j}) =
\argmax_{v_{j} \in V} P(v_{j}) P(a_{1} \ldots, a_n \,|\, v_{j}) \]
- Naive Bayes posteriors often unrealistically close to 1 or
0.
- If none of the training instances with target value $v_j$ have attribute
value $a_i$ Then
\[ \hat{P}(a_i\,|\,v_j) = 0 \text{, and...}\]
\[ \hat{P}(v_{j}) \prod_{i} \hat{P}(a_{i} \,|\, v_{j}) = 0 \]
Typical solution is Bayesian estimate for $\hat{P}(a_{i} \,|\, v_{j})$
\[ \hat{P}(a_{i} \,|\, v_{j}) \leftarrow \frac{n_{c} + mp}{n + m} \]
where
- $n$ is number of training examples for which $v=v_j$,
- $n_c$ number of examples for which $v=v_j$ and
$a=a_i$
- $p$ is prior estimate for $\hat{P}(a_{i} \,|\, v_{j})$
- $m$ is weight given to prior (i.e. number of ``virtual'' examples)
10 Learning to Classify Text
- Can we learn to classify which news articles are of
interest? which web pages are about astronomy?
- Naive Bayes is among most effective algorithms
- What attributes shall we use to represent text documents??
10.1 Text Attributes
- Target concept: IsItInteresting? : Document $\rightarrow
\{\oplus,\ominus\}$
- Represent each document by vector of words, one attribute
per word position in document, where $a_i$ is the $i^{th}$ word
in the document. For simplicity $doc \equiv a_1, a_2 \ldots$.
- Let $w_k$ be the $k$th word in the English vocabulary.
- Use training examples to estimate
\[P(\oplus), P(\ominus), P(doc\,|\,\oplus), P(doc\,|\,\ominus)\]
- Naive Bayes conditional independence assumption
\[ P(doc\,|\,v_j) = \prod_{i=1}^{length(doc)} P(a_i=w_k \,|\, v_j) \]
where $P(a_i=w_k\,|\, v_j)$ is probability that word in position $i$ is
$w_k$, given $v_j$.
- Also, assume that $\forall_{i,m \in \mathcal{N}} P(a_i=w_k\,|\,v_j) =
P(a_m=w_k\,|\,v_j)$, i.e., any word any place.
10.2 Learn Naive Bayes Text
- Given a set of Examples, and V.
- Collect all words and other tokens that occur in
Examples.
- $Vocabulary \leftarrow$ all distinct words and other tokens in $Examples$
- Calculate the required $P(v_{j})$ and $P(w_{k}\,|\,v_{j})$ probability terms.
- $docs_{j} \leftarrow $ subset of $Examples$ for which the target value is $v_{j}$
- $P(v_{j}) \leftarrow \frac{|docs_{j}|}{|Examples|}$
- $Text_{j} \leftarrow $ a single document created by
concatenating all members of $docs_{j}$
- $n \leftarrow$ total number of words in $Text_{j}$ (counting
duplicate words multiple times)
- for each word $w_{k}$ in $Vocabulary$
- $n_{k} \leftarrow$ number of times word $w_{k}$ occurs in
$Text_{j}$
- $P(w_{k}\,|\,v_{j}) \leftarrow \frac{n_{k} + 1}{n + |Vocabulary|}$
10.3 Classify Naive Bayes Text
- $positions \leftarrow$ all word positions in $Doc$ that contain
tokens found in $Vocabulary$
- Return $v_{NB}$, where
\[v_{NB} = \argmax_{v_{j} \in V} P(v_{j}) \prod_{i \in positions}P(a_{i}\,|\,v_{j}) \]
- This algorithm was shown to classify Usenet articles into
their appropriate newsgroups with 89% accuracy.
- A similar approach was proposed by Paul Graham in A Plan for Spam [4]. Several
implementations exist such as Spambayes [5].
11 Bayesian Belief Networks
- Naive Bayes assumption of conditional independence is too
restrictive, but learning is intractable without some such
assumptions. Hmmm.....
- Bayesian Belief networks describe conditional independence
among subsets of variables
- They allow combining prior knowledge about
(in)dependencies among variables with observed training data
- Also known as, Bayes nets.
11.1 Conditional Independence
- $X$ is conditionally independent of $Y$ given
$Z$ if the probability distribution governing $X$ is
independent of the value of $Y$ given the value of $Z$; that
is, if
\[ \forall_{x_i,y_j,z_k} P(X = x_i \,|\, Y = y_j, Z = z_k) = P(X = x_i \,|\, Z
= z_k) \] more compactly, we write \[ P(X \,|\, Y,Z) = P(X \,|\, Z) \]
- For example, $Thunder$ is conditionally independent of
$Rain$, given $Lightning$
\[ P(Thunder \,|\, Rain, Lightning) = P(Thunder \,|\, Lightning) \]
- Naive Bayes uses cond. indep. to justify
\[ \array{
P(X,Y\,|\,Z) &= & P(X\,|\,Y,Z)\cdot P(Y\,|\,Z) \\
&= & P(X\,|\,Z)\cdot P(Y\,|\,Z)
} \]
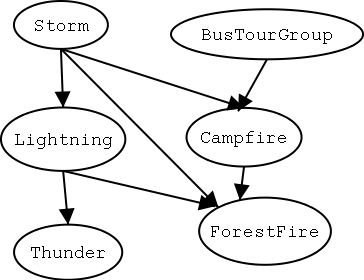
11.2 Bayesian Belief Network
- The network represents a set of conditional independence
assertions.
- Each node is asserted to be conditionally independent of its nondescendants, given its
immediate predecessors.
- It forms a directed acyclic graph.
- In general,
\[P(y_1, \ldots, y_n) = \prod_{i=1}^{n} P(y_i \,|\, Parents(Y_i)) \] where
$Parents(Y_i)$ denotes immediate predecessors of $Y_i$ in graph
- Joint distribution is fully defined by graph, plus the $P(y_i\,|\,Parents(Y_i))$.
11.3 Inference in Bayesian Networks
- How can one infer the (probabilities of) values of one or more network
variables, given observed values of others?
- Bayes net contains all information needed for this inference.
- If there is only one variable with an unknown value then easy to infer it.
- In the general case theproblem is NP hard.
- In practice, we can succeed in many cases:
- Exact inference methods work well for some network structures.
- Monte Carlo methods “simulate” the network randomly to calculate approximate solutions.
11.4 Learning Bayesian Networks
- Much of the research in Bayesian nets goes into how to use
them. We are, instead, interested in learning them.
- The network structure might be known or unknown.
- Training examples might provide values of all
network variables, or just some.
- If the structure known and we can observe all variables then its
easy as training a Naive Bayes classifier.
11.4.1 Learning Bayesian Networks
- Suppose structure known, variables partially
observable. For example we observe (ForestFire, Storm,
BusTourGroup, Thunder), but not (Lightning, Campfire)
- We can use a method similar to training neural network
with hidden units.
- Specifically, we will use gradient ascent.
- We want to converge to network $h$ that (locally) maximizes $P(D\,|\,h)$
11.4.2 Gradient Ascent for Bayes Nets
- Let $w_{ijk}$ denote one entry in the conditional probability table for
variable $Y_i$ in the network
\[ w_{ijk} = P(Y_i=y_{ij}\,|\,Parents(Y_i) = \text{the list} u_{ik} \text{of values}) \]
e.g., if $Y_i = Campfire$, then $u_{ik}$ might be $\langle Storm=T, BusTourGroup=F \rangle$
- Perform gradient ascent by repeatedly
- update all $w_{ijk}$ using training data $D$
\[w_{ijk} \leftarrow w_{ijk} + \eta \sum_{d \in D} \frac{P_h(y_{ij}, u_{ik} \,|\,
d)}{w_{ijk}} \]
- then, re-normalize the $w_{ijk}$ to assure $\sum_{j} w_{ijk} = 1$ and $0 \leq w_{ijk} \leq 1$
- If some of the probabilities are not present in the data they can be derived via inference from the net.
- Again, this only finds locally optimal conditional
probabilities.
11.5 The Expectation Maximization Algorithm
- The EM algorithm can also be used to learn a Bayes net.
- Calculate probabilities of unobserved variables, assuming
$h$.
- Calculate new $w_{ijk}$ to maximize $E[\ln P(D\,|\,h)]$ where $D$ now includes
both observed and (calculated probabilities of) unobserved variables.
- When the structure of the network is unknown we can use
algorithms that use greedy search to add/subtract edges and
nodes. This is an active area of research.
11.5.1 When To Use EM Algorithm
- Data is only partially observable.
- Unsupervised clustering (target value unobservable).
- Supervised learning (some instance attributes unobservable).
- Can be used to:
- Train Bayesian Belief Networks.
- Unsupervised clustering (AUTOCLASS).
- Learning Hidden Markov Models.
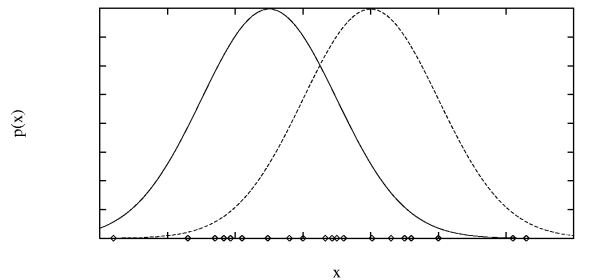
11.5.2 EM Example: Generating Data from k Gaussians
- Imagine the examples are generated by choosing instances
$x$ from $k$ Gaussians, with uniform probability.
- The learning task is to output a hypothesis that describes
the means $\langle \mu_1, \ldots, \mu_k \rangle$ of the $k$
distributions.
- We don't know which instance $x_i$ was generated by which Gaussian.
- We would like to find a maximum likelihood hypothesis for
those means, that is, the maximum likelihood estimates of
$\langle \mu_1, \ldots, \mu_k \rangle$.
- Think of full description of each instance as $y_i = \langle x_i, z_{i1}, z_{i2}
\rangle$, where
- $z_{ij}$ is 1 if $x_i$ generated by $j$th
Gaussian
- $x_i$ observable
- $z_{ij}$ unobservable
11.5.3 EM for Estimating k Means
- EM Algorithm: Pick random initial $h = \langle \mu_1, \mu_2 \rangle$, then iterate
- E step: Calculate the expected value $E[z_{ij}]$ of each hidden variable $z_{ij}$,
assuming the current hypothesis $h = \langle \mu_1, \mu_2 \rangle$ holds.
\[
\array{ E[z_{ij}] & = & \frac{p(x=x_i \,|\, \mu = \mu_j)}{\sum_{n=1}^{2} p(x = x_i \,|\, \mu=\mu_n)} \\
& = & \frac{e^{-\frac{1}{2 \sigma^2} (x_i -
\mu_j)^2}}{\sum_{n=1}^{2} e^{-\frac{1}{2 \sigma^2} (x_i - \mu_n)^2}}
} \]
- M step: Calculate a new maximum likelihood hypothesis $h' = \langle \mu_1', \mu_2'
\rangle$, assuming the value taken on by each hidden variable $z_{ij}$ is
its expected value $E[z_{ij}]$ calculated above. Replace $h =
\langle \mu_1, \mu_2 \rangle$ by $h' = \langle \mu_1', \mu_2' \rangle$.
\[ \mu_j \leftarrow \frac{\sum_{i=1}^m E[z_{ij}] x_i}{\sum_{i=1}^m E[z_{ij}]} \]
11.5.4 EM Algorithm
- Converges to local maximum likelihood $h$
and provides estimates of hidden variables $z_{ij}$
- In fact, local maximum in $E[\ln P(Y\,|\,h)]$
- $Y$ is complete (observable plus unobservable variables)
data
- Expected value is taken over possible values
of unobserved variables in $Y$
11.5.5 General EM Problem
Given:
- Observed data $X=\{x_1, \ldots, x_m\}$
- Unobserved data $Z=\{z_1, \ldots, z_m\}$
- Parameterized probability distribution $P(Y\,|\,h)$, where
- $Y=\{y_1, \ldots, y_m\}$ is the
full data $y_i = x_i \union z_i$ and $h$ are the parameters
Determine:
- $h$ that (locally) maximizes $E[\ln P(Y\,|\,h)]$
11.5.6 General EM Method
- Define likelihood function $Q(h' \,|\, h)$ which calculates $Y = X \union Z$ using
observed $X$ and current parameters $h$ to estimate $Z$
\[ Q(h' \,|\, h) \leftarrow E[ \ln P(Y \,|\, h') \,|\, h, X ] \]
- E step: Calculate $Q(h'\,|\,h)$ using the current hypothesis
$h$ and the observed data $X$ to estimate the probability distribution
over $Y$. \[ Q(h' \,|\, h) \leftarrow E[ \ln P(Y \,|\, h') \,|\, h, X ] \]
- M step: Replace hypothesis $h$ by the hypothesis $h'$
that maximizes this $Q$ function.
\[ h \leftarrow \argmax_{h'} Q(h' \,|\, h) \]
11.6 Summary of Bayes Nets
- Combine prior knowledge with observed data
- Impact of prior knowledge (when correct!) is to lower the
sample complexity
- Active research areas
- Extend from boolean to real-valued variables
- Parameterized distributions instead of tables
- Extend to first-order instead of propositional systems
- More effective inference methods
URLs
- Machine Learning book at Amazon, http://www.amazon.com/exec/obidos/ASIN/0070428077/multiagentcom/
- Slides by Tom Mitchell on Machine Learning, http://www-2.cs.cmu.edu/~tom/mlbook-chapter-slides.html
- Wikipedia:Bayes Theorem, http://www.wikipedia.org/wiki/Bayes_theorem
- A Plan for Spam by Paul Graham, http://www.paulgraham.com/spam.html
- Spambayes Homepage, http://spambayes.sourceforge.net/
This talk available at http://jmvidal.cse.sc.edu/talks/bayesianlearning/
Copyright © 2009 José M. Vidal
.
All rights reserved.
25 February 2003, 02:16PM